Abstract
Background:
Functional microRNAs (miRNAs) in exosomes have been recognised as potential stable biomarkers in cancers. The aim of this study is to identify specific miRNAs in exosome as serum biomarkers for the early detection of recurrence in human colorectal cancer (CRC).
Methods:
Serum samples were sequentially obtained from six patients with and without recurrent CRC. The miRNAs were purified from exosomes, and miRNA microarray analysis was performed. The miRNA expression profiles and copy number aberrations were explored using microarray and array CGH analyses in 124 CRC tissues. Then, we validated exosomal miRNAs in 2 serum sample sets (90 and 209 CRC patients) by quantitative real-time RT–PCR.
Results:
Exosomal miR-17-92a cluster expression level in serum was correlated with the recurrence of CRC. Exosomal miR-19a expression levels in serum were significantly increased in patients with CRC as compared with healthy individuals with gene amplification. The CRC patients with high exosomal miR-19a expression showed poorer prognoses than the low expression group (P<0.001).
Conclusions:
Abundant expression of exosomal miR-19a in serum was identified as a prognostic biomarker for recurrence in CRC patients.
Similar content being viewed by others
Main
Colorectal cancer (CRC) is the second most common cause of cancer mortality worldwide (Soerjomataram et al, 2012). Despite surgical resection and pre- and postoperative chemotherapy and molecular targeting therapy, relapse is common in patients with CRC (Akiyoshi et al, 2012; Amano et al, 2014). Recurrence, particularly liver metastasis, clearly contributes to the survival of patients with CRC. Many studies have examined potential biomarkers of recurrence in CRC in an attempt to improve patients’ prognosis, and microRNAs (miRNAs), the small noncoding RNAs that are associated with the development of cancer, have been shown to be potential biomarkers in various types of cancers (Calin and Croce, 2006; Cortez et al, 2012; Komatsu et al, 2014).
Exosomes secreted from sheep reticulocytes were first reported in1983 (Pan and Johnstone, 1983), and the mechanism of exosome-mediated miRNA transfer was verified in 2007 (Valadi et al, 2007). Exosomes have the capacity to envelop specific miRNAs, maintain the integrity of contents (such as nucleic acids) in circulation and function as a tool for invasive access while reflecting the condition of donor cells (Palma et al, 2012; Ge et al, 2014; Kowal et al, 2014). Through these mechanisms, exosomal contents can be transmitted from donor cells to recipient cells. Thus, exosomal miRNAs can contribute to cancer development. Recent studies have shown that detection of specific exosomes may represent a novel diagnostic tool; indeed, ongoing studies are investigating the application of exosome contents as biomarkers and capsules for delivery of therapeutic drugs (van der Meel et al, 2014; Yoshioka et al, 2014b). Thus, exosomal miRNAs may represent both biomarkers and potential therapeutic targets (Hoshino and Matsubara, 2013). In terms of exosomal function, exosome-mediated cell-to-cell communication may play an important role in the development of cancer. Several reports have shown that specific exosomal miRNAs are related to the phenotypes of certain types of cancer and therefore may function as meaningful biomarkers (Tanaka et al, 2013; Ogata-Kawata et al, 2014; Xue et al, 2014). We previously revealed that a specific exosomal miRNA is a biomarker to predict the recurrence of liver cancer (Sugimachi et al, 2015). Moreover, many studies have investigated the application of miRNAs as molecular therapies in the treatment of CRC (Hoshino and Matsubara, 2013; Li and Rana, 2014).
Distant metastasis mediated through the hematogenous route directly contributes to the poor prognosis in patients with CRC (Field and Lipton, 2007; (Amano et al, 2014). In addition, CRC cells secrete exosomes into the circulation. Therefore, the aim of this study was to identify serum exosomal miRNAs as biomarkers of recurrence in human CRC. Among the miRNAs that have been shown to be associated with CRC, we focussed on miRNAs with copy number aberrations as evaluated by array comparative genomic CGH (aCGH) in this study because these miRNAs should be robust, stable molecular targets. Finally, we examined the expression of exosomal miRNAs using microarray and quantitative real-time reverse transcription–PCR (qRT–PCR).
Materials and Methods
Patients and sample collection
All serum and tissue samples were obtained from patients who underwent primary tumour resection at Kyushu University Beppu Hospital and affiliated hospitals between 1992 and 2007. Written informed consent was obtained from all patients. The study was approved by the institutional review board, the Ethics and Indications Committee of Kyushu University. We used a total of 227 serum samples from CRC patients, of which 6 samples were used for microarray and 221 samples were used for qRT-PCR. Serum samples were also obtained from 28 healthy volunteers for qRT–PCR. A total of 8 tumour tissues and 4 nearby normal tissues from CRC patients were used for aCGH, copy number analysis and miR microarray. All serum samples were redundant blood obtained for the purpose of examination during preoperative and follow-up phases. All patients had a histological diagnosis of CRC and were closely followed up every 3 months. The average overall survival time after resection was 4.64 years in the 209 patients from which samples were used for qRT–PCR. Tumour recurrence was observed in 132 of 165 patients with average interval of 4.82 years. We treated patients in accordance with the Japanese Society of Cancer of the Colon and Rectum Guidelines for the Treatment of Colorectal Cancer. Tumour tissue and normal tissue were immediately cut from the resected colorectal tissue and placed in RNAlater (Ambion, Palo Alto, CA, USA) or embedded in Tissue Tek OCT medium (Sakura, Tokyo, Japan), frozen in liquid nitrogen and kept at −80 °C until RNA extraction. Clinicopathological factors and clinical stage were classified using the TNM system of classification. All data for the samples, including patient age, gender, tumour size and depth, lymphatic invasion, lymph node metastasis, vascular invasion, liver metastasis, peritoneal dissemination, distant metastasis, clinical stage and histological grade, were obtained from the clinical and pathological records.
Isolation of exosomes from serum
For the ultracentrifuge methods, serum of 1.5 ml for microarray and 1.0 ml for qRT–PCR were ultracentrifuged at 100 000 g for 70 min at 4 °C. The pellets were washed with 11 ml of phosphate-buffered saline as described elsewhere (Sugimachi et al, 2015). Exosomes were extracted from a very small volume of serum using a Total Exosome Isolation Kit (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. The serum volume was fixed at 100 μl for validation by qRT–PCR of samples from 90 patients with CRC and 12 healthy volunteers.
Transmission electron microscopy
The isolated exosomes fraction was dissolved in HEPES buffer, and a drop of the suspension was placed on a sheet. A carbon-coated copper grid was floated on the drop for 10 s. The grid was then removed, and excess liquid was drained from the edge of the grid using a piece of clean filter paper. The grid was touched onto a drop of 2% uranyl acetate or phosphotungstic acid, pH 7.0, for ∼5 s, and excess liquid was removed. The grid was allowed to dry for several minutes and was then examined using a JEM1400 microscope (JEOL, Akishima, Japan) (Tanaka et al, 2013).
Extraction of total RNA and miRNA
Frozen tissue specimens were homogenised, and total RNA was extracted using a modified acid-guanidine-phenol-chloroform method. The RNA was extracted from exosomes in serum, using miRNeasy mini kit (Qiagen, Venlo, Netherlands) according to the manufacture’s protocol. Exosomes purified from a certain volume of serum were diluted with 1 ml of Qiazol Lysis Reagent (Qiagen). After 5 min of incubation at room temperature, 10 μl of 0.1 nM cel-miR-39 mimic was added to each sample followed by vortexing for 30 s. Subsequent extraction and cartridge work were carried out according to the manufacturer’s protocol (Kosaka et al, 2010) (Yamamoto et al, 2011).
miRNA microarray analysis
The quality of total RNAs from the exosome fraction was evaluated using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). No DNA contamination was observed. Exosomal miRNA expression profiles were examined using a miRNA microarray analysis system (3D-Gene, Toray, Tokyo, Japan).
Twelve samples (8 from cancer tissues and 4 from normal epithelia) were selected and used for the microarray. The concentration and purity of the total RNAs were assessed using a spectrophotometer, and RNA integrity was verified using an Agilent 2100 Bioanalyzer. Total RNA (100 ng) was directly labelled with cyanine3-CTP (Cy3), without fractionation or amplification, using an Agilent protocol that produces precise and accurate measurements spanning a linear dynamic range from 0.2 amol to 2 fmol of input miRNA. Total RNAs (50 ng) from 12 samples were competitively hybridised to a miRNA array (Agilent Microarray) according to the manufacturer’s protocol. A list of miRNAs contained in the array is available from version 14.0 of the Sanger miRNA database (http://microrna.sanger.ac.uk). For data analysis, the intensity of each hybridisation signal was evaluated using Feature Extraction Software (Agilent Technologies) that examines multiple probes and multiple features per probe and studies the measurements and errors for each miRNA. The observed values were imported into GeneSpring GX version 11.0 (Agilent Technologies). Generated miRNA profiles were normalised to the amount of input total RNA.
aCGH and copy number analysis
A total of 124 colorectal tissues were prepared for aCGH analysis. For genome profiling, an Agilent Human Genome Microarray Kit 244K (Agilent Technologies) was used according to the manufacturer’s instructions. The raw copy number data for each sample provided by aCGH were analysed using GISTIC algorithm (Mermel et al, 2011).
qRT–PCR analysis of miRNAs
Exosomal miRNA expression was assayed using qRT–PCR in serum sample of CRC patients. The cDNA was synthesised from total RNA using Taqman MicroRNA primers specific for hsa-miR-16a, hsa-miR-17a, hsa-miR-18a, hsa-miR-19a, hsa-miR-19b, has-miR-20a and hsa-miR-92a (Applied Biosystems, Tokyo, Japan) and a Taqman Micro-RNA Reverse Transcription Kit (Applied Biosystems). Relative quantification of miRNA expression was calculated using the 2-ΔΔCt method. The miR-16a was used as an internal control because it has been reported to be a reliable endogenous control for analysis of miRNA by RT–PCR in humans (Davoren et al, 2008). The raw data were presented as the relative quantity of target miRNA, normalised with respect to miR-16a and compared with a reference sample. Real-time monitoring of PCR products of samples from 90 patients with CRC and 12 healthy volunteers was performed using an ANI PRISM 7300 (Applied Biosystems) and Taqman Universal PCR Master Mix (Applied Biosystems) following the manufacturer’s protocol. Real-time monitoring of PCR products of samples from 209 patients with CRC and 16 healthy volunteers was performed using LightCycler480 (Roche Applied Science, Basel, Switzerland) and Taqman Universal PCR Master Mix (Applied Biosystems) following the manufacturer’s protocol. The amplification protocol included an initial denaturation step at 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 60 s (Yoshioka et al, 2014a).
Statistical analysis
For continuous variables, data were expressed as mean±s.d. Survival curves were plotted according to the Kaplan–Meier method and the generalised log-rank test was applied to compare the survival curves. Differences between groups were estimated using the χ2 test, Student’s t-test, repeated-measures analysis of variance (ANOVA), log-rank test and Cox regression model. All tests were analysed using JMP 5 software (JMP, Cary, NC, USA), and the findings were considered significant when the P-value was <0.05.
Results
Identification of exosomal RNA in serum
To verify our ultracentrifugation method for isolation of exosomes, we examined exosome using transmission electron microscopy. We captured images of round microvesicles having diameters of ∼50 nm in the exosome-rich fraction (Supplementary Figure 1A and B). Exosomal RNA was purified and miRNAs were observed as products of ∼25 nucleotides in length (Supplementary Figure 1C).
Relationship between serum exosomal miRNAs and the progression of CRC
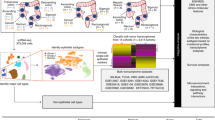
The schematic explanation of procedure to select exosomal miRNAs associated with CRC recurrence is shown in Supplementary Figure 2. We examined the relationship between serum exosomal miRNAs and CRC progression using four serum samples that were sequentially obtained from a CRC patient with recurrence of liver metastasis during pre- and postoperative phases of primary resection and pre- and postoperative phases of hepatectomy as well as two serum samples that were sequentially obtained from a CRC patient without recurrence during pre- and postoperative phases of primary resection. Exosomal RNA extracted from each serum sample was used for microarray profiling. Eighteen exosomal microRNAs were increased and 46 exosomal miRNAs were decreased by more than two-fold in serum from the patient with liver metastasis as compared with the patient without recurrence (Table 1). Interestingly, the expressions of 6 exosomal miRNAs (i.e., miR-19a, miR-19b, miR-23a, miR-92a, miR-320a and miR-4437) were synchronised with the development of liver metastasis. Therefore, these six miRNAs were candidate miRNAs associated with liver metastasis in CRC. Among the six miRNAs, miR-19a and miR-92a were significantly upregulated in CTC patients compared with healthy control, and were therefore further evaluated.
miRNA cluster exhibited genomic amplification in CRC tissue
Of the 417 miRNAs analysed, changes in the expression of 58 miRNAs were shown to be related to alternations in a genomic locus. Twenty-four miRNAs had correlation coefficients of over 0.5, and hsa-miR-25-106b and hsa-miR-17-92a clusters were identified (Table 2).
Expression of the exosomal miR-17-92a cluster in serum
Three of six serum exosomal miRNAs exhibiting expression correlating with CRC in microarray analysis were located within the miR-17-92a locus. As mentioned above, mir-17-92a clusters were upregulated by genetic amplification in CRC tissue. Therefore, we selected miRNAs within the miR-17-92a cluster as candidate miRNAs associated with CRC. Expression of these six exosomal miRNAs (i.e., miR-17, miR-18a, miR-19a, miR-19b, miR20a and miR-92a) in serum was assessed by qRT–PCR in 102 serum samples from 90 patients with CRC and 12 healthy volunteers. Of the six serum exosomal miRs analysed, miR-19a and miR-92a were significantly increased in CRC patients compared with healthy volunteers (Figure 1A), whereas the other four miRs did not show significant difference. Then, the expression of exosomal miR-19a and miR-92a in each stage of CRC patient was explored (Figures 1B and C). Interestingly, those exosomal miRNAs were upregulated in both early and advanced stages of CRC compared with healthy controls.
Expression of six microRNAs in the miR-17-92 cluster. Quantitative RT–PCR using Taqman miRNA assays was used to investigate the expression of the six miRNAs in exosomes purified from serum. The obtained values were normalised to hsa-miR-16a as an internal control. (A) Expression of 6 serum exosomal miRNAs in 6 healthy volunteers and 90 patients with CRC. (B) Expression of serum exosomal miR-19a in healthy individuals and patients with different stages of CRC. (C) Expression of serum exosomal miR-92a in healthy individuals and patients with different stages of CRC.
Expression of exosomal miR-19a and clinicopathological characteristics
For clinicopathological analysis, we classified the 209 CRC serum samples into 2 groups using the average of miR-19a expression level as determined from 16 healthy volunteers. Patients in the high exosomal miR-19a expression group (n=133) had more frequent occurrence of malignancy-related factors, including serosal invasion (P=0.0224), lymphatic permeation (P=0.0028), lymph node metastasis (P=0.0206), liver metastasis (P=0.0486) and higher TNM stage (P=0.0005), as compared with the low exosomal miR-19a expression group (n=76; Table 3).
Prognostic significance of exosomal miR-19a expression
Using the two exosomal miR-19a expression groups described in the previous section, we analysed the association between miR-19a expression and survival rates. We found that high exosomal miR-19a expression was significantly associated with poorer survival as compared with low exosomal miR-19a expression (P=0.0004; Figure 2A). In 164 samples from patients with CRC (excluding stage IV), disease-free survival rate were significantly lower in patients with high exosomal miR-19a expression than in patients with low exosomal miR-19a expression (P=0.0002; Figure 2B). In addition, univariate and multivariate analysis showed that serum exosomal miR-19a expression was an independent risk factor for overall survival (Table 4a) and disease-free survival (Table 4b) in CRC patients.
Kaplan–Meier survival curves for CRC patients classified according to the miR-19a expression level. (A) Overall survival curve of 209 patients with CRC. Two groups were divided according to the average exosomal miR-19a expression level in serum of healthy individual. Patients with high expression of exosomal miR-19a in serum had significantly poorer prognoses than patients with low expression of exosomal miR-19a. (B) Disease-free survival curve for 164 patients with CRC.
Discussion
The aim of this study was to identify specific serum exosomal miRNAs as biomarkers reflecting the progression of CRC. We hypothesised that CRC cells secreted exosomes containing miRNAs into the blood in order to transport signals to recipient cells. Our findings indicate that serum exosomal miR-19a could be a potential biomarker to predict recurrence of CRC.
In this study, analysis of serum exosomal miRNA expression profiles revealed that six6 serum exosomal miRNAs were regulated concordant with CRC progression. We confirmed that the expression of serum exosomal miR-19a was higher in patients with CRC than in healthy controls using two different sets of serum samples. Interestingly, regulation of miRNA expression has been shown to be dependent on the location of miRNAs within cancer-associated loci that exhibit genomic alternations, epigenetic regulation and abnormalities of essential proteins involved in miRNA maturation (Calin and Croce, 2006). Genomic alternations have been shown to cause abnormal miRNA expression. In addition, hsa-miR-25-106b and hsa-miR-17-92a clusters have cancer-promoting function (Tan et al, 2014). In particular, the hsa-miR-17-92a, a well-known oncogenic miRNAs cluster, has been shown to be upregulated in various types of cancers, such as of colon cancer, lung cancer and lymphoma (Hayashita et al, 2005; Diosdado et al, 2009; Olive et al, 2010). These findings supported our hypothesis that CRC cells secrete exosomal miRNAs to survive. Cells overexpressing miR-19a because of genomic alternations may secrete miR-19a-rich exosomes containing miR-19a. It has been reported that miR-19a promotes proliferation and invasion of cancer cells by gain- or loss-of-function experiments (Zhang et al, 2012; Feng et al, 2014). The downstream signalling pathway of miR-19a has not been clarified yet, but several reports have shown that miR-19a plays an oncogenic role by repressing tumour suppressor gene PTEN in lymphoma cells, gastric cancer cells and bladder cancer cells (Olive et al, 2009; Liang et al, 2011; Wang et al, 2013; Feng et al, 2014). We showed here that high expression of exosomal miR-19a was associated with poor prognosis in patients with CRC, suggesting that exosomal miR-19a may be a potential marker of poor prognosis in CRC. Indeed, in our data set, we found that high levels of exosomal miR-19a in serum of CRC patients were associated with tumour recurrence. Interestingly, miR-19a increased in patients with early-stage CRC as well as those with advanced stage. Those results were concordant with the previous study that reported serum miRNAs including miR-19a could be a biomarker for early detection of CRC (Zheng et al, 2014). Those results suggested that miR-19a might function at early stage of colorectal oncogenesis. Recently, Zheng et al (2014) reported that a four-miRNA panel (miR-19a-3p, miR-223-3p, miR-92a-3p and miR-422a) for serum samples could successfully differentiate CRC from colon adenoma and healthy control. Those data were consist with our results that miR-19a and miR-92a were significantly upregulated in serum of CRC patients, and our present data even proved that those serum miRs were functional secreted in exosome.
We observed a relationship between miR-19a expression and CRC progression, but the mechanisms through which cancer cells secreted exosome-containing specific miRNAs have been unknown in detail. Further studies are required to clarify the relationship between serum exosomal miR-19a and intercellular signalling in CRC.
In conclusion, serum exosomal miR-19a was upregulated in CRC independent of clinical staging. Serum exosomal miR-19a may be a potential marker of prognosis and a therapeutic target in CRC.
Change history
14 July 2015
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Akiyoshi T, Kobunai T, Watanabe T (2012) Recent approaches to identifying biomarkers for high-risk stage II colon cancer. Surg Today 42 (11): 1037–1045.
Amano R, Yamada N, Nakata B, Kimura K, Yashiro M, Ohira M, Hirakawa K (2014) Prognostic indicator for the resection of liver metastasis of colorectal cancer. Surg Today 44 (7): 1287–1292.
Calin GA, Croce CM (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6 (11): 857–866.
Cortez MA, Welsh JW, Calin GA (2012) Circulating microRNAs as noninvasive biomarkers in breast cancer. Recent Results Cancer Res 195: 151–161.
Davoren PA, McNeill RE, Lowery AJ, Kerin MJ, Miller N (2008) Identification of suitable endogenous control genes for microRNA gene expression analysis in human breast cancer. BMC Mol Biol 9: 76.
Diosdado B, van de Wiel MA, Terhaar Sive Droste JS, Mongera S, Postma C, Meijerink WJ, Carvalho B, Meijer GA (2009) MiR-17-92 cluster is associated with 13q gain and c-myc expression during colorectal adenoma to adenocarcinoma progression. Br J Cancer 101 (4): 707–714.
Feng Y, Liu J, Kang Y, He Y, Liang B, Yang P, Yu Z (2014) miR-19a acts as an oncogenic microRNA and is up-regulated in bladder cancer. J Exp Clin Cancer Res 33: 67.
Field K, Lipton L (2007) Metastatic colorectal cancer-past, progress and future. World J Gastroenterol 13 (28): 3806–3815.
Ge Q, Zhou Y, Lu J, Bai Y, Xie X, Lu Z (2014) miRNA in plasma exosome is stable under different storage conditions. Molecules 19 (2): 1568–1575.
Hayashita Y, Osada H, Tatematsu Y, Yamada H, Yanagisawa K, Tomida S, Yatabe Y, Kawahara K, Sekido Y, Takahashi T (2005) A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res 65 (21): 9628–9632.
Hoshino I, Matsubara H (2013) MicroRNAs in cancer diagnosis and therapy: from bench to bedside. Surg Today 43 (5): 467–478.
Komatsu S, Ichikawa D, Hirajima S, Kawaguchi T, Miyamae M, Okajima W, Ohashi T, Arita T, Konishi H, Shiozaki A, Fujiwara H, Okamoto K, Yagi N, Otsuji E (2014) Plasma microRNA profiles: identification of miR-25 as a novel diagnostic and monitoring biomarker in oesophageal squamous cell carcinoma. Br J Cancer 111 (8): 1614–1624.
Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, Ochiya T (2010) Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem 285 (23): 17442–17452.
Kowal J, Tkach M, Thery C (2014) Biogenesis and secretion of exosomes. Curr Opin Cell Biol 29: 116–125.
Li Z, Rana TM (2014) Therapeutic targeting of microRNAs: current status and future challenges. Nat Rev Drug Discov 13 (8): 622–638.
Liang Z, Li Y, Huang K, Wagar N, Shim H (2011) Regulation of miR-19 to breast cancer chemoresistance through targeting PTEN. Pharm Res 28 (12): 3091–3100.
Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G (2011) GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol 12: R41.
Ogata-Kawata H, Izumiya M, Kurioka D, Honma Y, Yamada Y, Furuta K, Gunji T, Ohta H, Okamoto H, Sonoda H, Watanabe M, Nakagama H, Yokota J, Kohno T, Tsuchiya N (2014) Circulating exosomal microRNAs as biomarkers of colon cancer. PLoS One 9 (4): e92921.
Olive V, Bennett MJ, Walker JC, Ma C, Jiang I, Cordon-Cardo C, Li QJ, Lowe SW, Hannon GJ, He L (2009) miR-19 is a key oncogenic component of mir-17-92. Genes Dev 23 (24): 2839–2849.
Olive V, Jiang I, He L (2010) mir-17-92, a cluster of miRNAs in the midst of the cancer network. Int J Biochem Cell Biol 42 (8): 1348–1354.
Palma J, Yaddanapudi SC, Pigati L, Havens MA, Jeong S, Weiner GA, Weimer KM, Stern B, Hastings ML, Duelli DM (2012) MicroRNAs are exported from malignant cells in customized particles. Nucleic Acids Res 40 (18): 9125–9138.
Pan BT, Johnstone RM (1983) Fate of the transferrin receptor during maturation of sheep reticulocytes in vitro: selective externalization of the receptor. Cell 33 (3): 967–978.
Soerjomataram I, Lortet-Tieulent J, Parkin DM, Ferlay J, Mathers C, Forman D, Bray F (2012) Global burden of cancer in 2008: a systematic analysis of disability-adjusted life-years in 12 world regions. Lancet 380 (9856): 1840–1850.
Sugimachi K, Matsumura T, Hirata H, Uchi R, Ueda M, Ueo H, Shinden Y, Iguchi T, Eguchi H, Shirabe K, Ochiya T, Maehara Y, Mimori K (2015) Identification of a bona fide microRNA biomarker in serum exosomes that predicts hepatocellular carcinoma recurrence after liver transplantation. Br J Cancer 112 (3): 532–538.
Tan W, Li Y, Lim SG, Tan TM (2014) miR-106b-25/miR-17-92 clusters: polycistrons with oncogenic roles in hepatocellular carcinoma. World J Gastroenterol 20 (20): 5962–5972.
Tanaka Y, Kamohara H, Kinoshita K, Kurashige J, Ishimoto T, Iwatsuki M, Watanabe M, Baba H (2013) Clinical impact of serum exosomal microRNA-21 as a clinical biomarker in human esophageal squamous cell carcinoma. Cancer 119 (6): 1159–1167.
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9 (6): 654–659.
van der Meel R, Fens MH, Vader P, van Solinge WW, Eniola-Adefeso O, Schiffelers RM (2014) Extracellular vesicles as drug delivery systems: lessons from the liposome field. J Control Release 195: 72–85.
Wang F, Li T, Zhang B, Li H, Wu Q, Yang L, Nie Y, Wu K, Shi Y, Fan D (2013) MicroRNA-19a/b regulates multidrug resistance in human gastric cancer cells by targeting PTEN. Biochem Biophys Res Commun 434 (3): 688–694.
Xue W, Dahlman JE, Tammela T, Khan OF, Sood S, Dave A, Cai W, Chirino LM, Yang GR, Bronson R, Crowley DG, Sahay G, Schroeder A, Langer R, Anderson DG, Jacks T (2014) Small RNA combination therapy for lung cancer. Proc Natl Acad Sci USA 111 (34): E3553–E3561.
Yamamoto Y, Yoshioka Y, Minoura K, Takahashi RU, Takeshita F, Taya T, Horii R, Fukuoka Y, Kato T, Kosaka N, Ochiya T (2011) An integrative genomic analysis revealed the relevance of microRNA and gene expression for drug-resistance in human breast cancer cells. Mol Cancer 10: 135.
Yoshioka Y, Kosaka N, Konishi Y, Ohta H, Okamoto H, Sonoda H, Nonaka R, Yamamoto H, Ishii H, Mori M, Furuta K, Nakajima T, Hayashi H, Sugisaki H, Higashimoto H, Kato T, Takeshita F, Ochiya T (2014a) Ultra-sensitive liquid biopsy of circulating extracellular vesicles using ExoScreen. Nat Commun 5: 3591.
Yoshioka Y, Uehara K, Ebata T, Yokoyama Y, Mitsuma A, Ando Y, Nagino M (2014b) Postoperative complications following neoadjuvant bevacizumab treatment for advanced colorectal cancer. Surg Today 44 (7): 1300–1306.
Zhang J, Xiao Z, Lai D, Sun J, He C, Chu Z, Ye H, Chen S, Wang J (2012) miR-21, miR-17 and miR-19a induced by phosphatase of regenerating liver-3 promote the proliferation and metastasis of colon cancer. Br J Cancer 107 (2): 352–359.
Zheng G, Du L, Yang X, Zhang X, Wang L, Yang Y, Li J, Wang C (2014) Serum microRNA panel as biomarkers for early diagnosis of colorectal adenocarcinoma. Br J Cancer 111 (10): 1985–1992.
Acknowledgements
We thank K Oda, M. Kasagi, S Kono, T Kawano and M Aoyagi for their technical assistance. This work was supported in part by the following grants and foundations: Japan Science and Technology Agency (JST); Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Science Research (Grant Numbers 25430111, 25461953, 25861199, 25861200 and 26861085); Japan Science and Technology Agency (JSTA) A-step, Grant Number AS242Z03987P); the Founding Program for Next Generation World-Leading Researchers (LS094) and NEDO.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Matsumura, T., Sugimachi, K., Iinuma, H. et al. Exosomal microRNA in serum is a novel biomarker of recurrence in human colorectal cancer. Br J Cancer 113, 275–281 (2015). https://doi.org/10.1038/bjc.2015.201
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2015.201
Keywords
This article is cited by
-
Circulating exosomal microRNAs as potential prognostic biomarkers in gastrointestinal cancers: a systematic review and meta-analysis
Cancer Cell International (2023)
-
Recent advances of small extracellular vesicle biomarkers in breast cancer diagnosis and prognosis
Molecular Cancer (2023)
-
Identification of exosomal microRNA panel as diagnostic and prognostic biomarker for small cell lung cancer
Biomarker Research (2023)
-
Migrasomes, a new mode of intercellular communication
Cell Communication and Signaling (2023)
-
Rebound increase in microRNA levels at the end of 5-FU-based therapy in colorectal cancer patients
Scientific Reports (2023)